Note
Go to the end to download the full example code.
Petrophysically guided inversion: Joint linear example with nonlinear relationships#
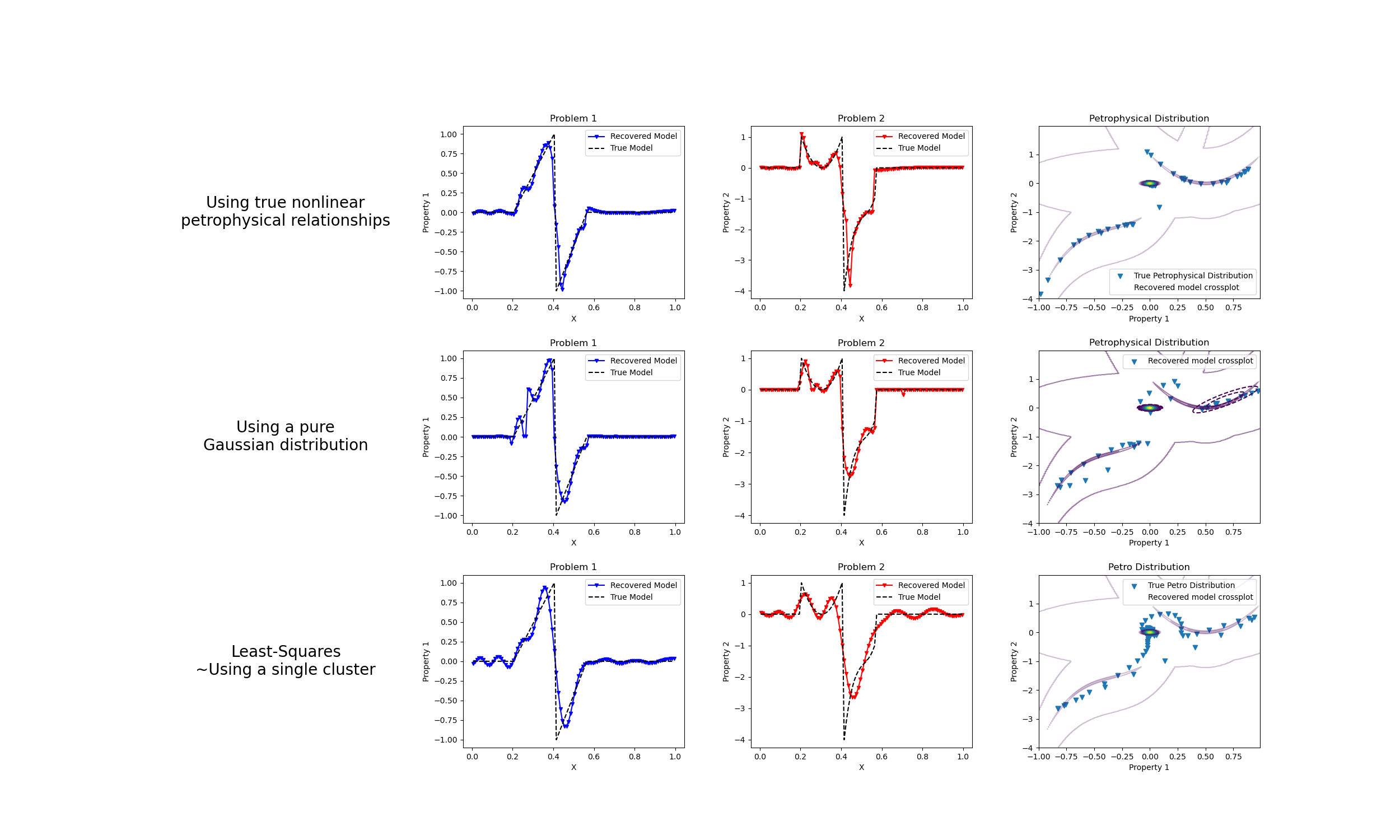
We do a comparison between the classic least-squares inversion and our formulation of a petrophysically guided inversion. We explore it through coupling two linear problems whose respective physical properties are linked by polynomial relationships that change between rock units.
Running inversion with SimPEG v0.23.1.dev10+gf697d2455
simpeg.InvProblem will set Regularization.reference_model to m0.
simpeg.InvProblem will set Regularization.reference_model to m0.
simpeg.InvProblem will set Regularization.reference_model to m0.
simpeg.InvProblem is setting bfgsH0 to the inverse of the eval2Deriv.
***Done using the default solver Pardiso and no solver_opts.***
Alpha scales: [np.float64(3.4917129103789835), np.float64(0.0), np.float64(3.490067899032831e-06), np.float64(0.0)]
Calculating the scaling parameter.
Scale Multipliers: [0.10209669 0.89790331]
<class 'simpeg.regularization.pgi.PGIsmallness'>
Initial data misfit scales: [0.10209669 0.89790331]
model has any nan: 0
=============================== Projected GNCG ===============================
# beta phi_d phi_m f |proj(x-g)-x| LS Comment
-----------------------------------------------------------------------------
x0 has any nan: 0
0 1.94e+01 3.00e+05 0.00e+00 3.00e+05 1.41e+02 0
geophys. misfits: 956.1 (target 30.0 [False]); 98.4 (target 30.0 [False]) | smallness misfit: 2974.7 (target: 200.0 [False])
Beta cooling evaluation: progress: [956.1 98.4]; minimum progress targets: [240000. 240000.]
1 1.94e+01 1.86e+02 4.16e+01 9.92e+02 9.81e+01 0
geophys. misfits: 426.7 (target 30.0 [False]); 38.8 (target 30.0 [False]) | smallness misfit: 1374.6 (target: 200.0 [False])
Beta cooling evaluation: progress: [426.7 38.8]; minimum progress targets: [764.9 78.7]
2 1.94e+01 7.84e+01 4.04e+01 8.63e+02 3.36e+01 0 Skip BFGS
geophys. misfits: 416.5 (target 30.0 [False]); 38.5 (target 30.0 [False]) | smallness misfit: 1361.9 (target: 200.0 [False])
Beta cooling evaluation: progress: [416.5 38.5]; minimum progress targets: [341.3 31. ]
Decreasing beta to counter data misfit decrase plateau.
3 9.70e+00 7.71e+01 4.05e+01 4.70e+02 8.91e+01 0 Skip BFGS
geophys. misfits: 147.5 (target 30.0 [False]); 30.8 (target 30.0 [False]) | smallness misfit: 1459.8 (target: 200.0 [False])
Beta cooling evaluation: progress: [147.5 30.8]; minimum progress targets: [333.2 30.8]
4 9.70e+00 4.27e+01 4.29e+01 4.59e+02 7.66e+00 0
geophys. misfits: 144.6 (target 30.0 [False]); 30.7 (target 30.0 [False]) | smallness misfit: 1445.4 (target: 200.0 [False])
Beta cooling evaluation: progress: [144.6 30.7]; minimum progress targets: [118. 30.]
Decreasing beta to counter data misfit decrase plateau.
5 4.85e+00 4.23e+01 4.29e+01 2.51e+02 7.24e+01 0 Skip BFGS
geophys. misfits: 71.7 (target 30.0 [False]); 26.3 (target 30.0 [True]) | smallness misfit: 1649.3 (target: 200.0 [False])
Beta cooling evaluation: progress: [71.7 26.3]; minimum progress targets: [115.7 30. ]
Updating scaling for data misfits by 1.140444791511067
New scales: [0.11478968 0.88521032]
6 4.85e+00 3.15e+01 4.46e+01 2.48e+02 6.03e+01 0
geophys. misfits: 59.7 (target 30.0 [False]); 26.4 (target 30.0 [True]) | smallness misfit: 1606.0 (target: 200.0 [False])
Beta cooling evaluation: progress: [59.7 26.4]; minimum progress targets: [57.3 30. ]
Decreasing beta to counter data misfit decrase plateau.
Updating scaling for data misfits by 1.135983417110108
New scales: [0.12839499 0.87160501]
7 2.43e+00 3.07e+01 4.46e+01 1.39e+02 8.75e+01 0
geophys. misfits: 22.6 (target 30.0 [True]); 21.5 (target 30.0 [True]) | smallness misfit: 2066.3 (target: 200.0 [False])
Beta cooling evaluation: progress: [22.6 21.5]; minimum progress targets: [47.7 30. ]
Warming alpha_pgi to favor clustering: 1.3628383796940546
8 2.43e+00 2.16e+01 4.84e+01 1.39e+02 8.66e+01 0
geophys. misfits: 20.5 (target 30.0 [True]); 23.0 (target 30.0 [True]) | smallness misfit: 1686.4 (target: 200.0 [False])
Beta cooling evaluation: progress: [20.5 23. ]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 1.886068723148174
9 2.43e+00 2.27e+01 4.94e+01 1.43e+02 3.70e+01 0
geophys. misfits: 16.9 (target 30.0 [True]); 24.2 (target 30.0 [True]) | smallness misfit: 1450.4 (target: 200.0 [False])
Beta cooling evaluation: progress: [16.9 24.2]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 2.8421507966954223
10 2.43e+00 2.33e+01 5.19e+01 1.49e+02 2.78e+01 0
geophys. misfits: 14.2 (target 30.0 [True]); 26.0 (target 30.0 [True]) | smallness misfit: 1178.1 (target: 200.0 [False])
Beta cooling evaluation: progress: [14.2 26. ]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 4.638518468807684
11 2.43e+00 2.44e+01 5.56e+01 1.59e+02 5.97e+01 0
geophys. misfits: 13.6 (target 30.0 [True]); 28.0 (target 30.0 [True]) | smallness misfit: 914.1 (target: 200.0 [False])
Beta cooling evaluation: progress: [13.6 28. ]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 7.58425007658527
12 2.43e+00 2.61e+01 6.00e+01 1.72e+02 9.52e+01 0
geophys. misfits: 14.8 (target 30.0 [True]); 30.3 (target 30.0 [False]) | smallness misfit: 702.0 (target: 200.0 [False])
Beta cooling evaluation: progress: [14.8 30.3]; minimum progress targets: [30. 30.]
Decreasing beta to counter data misfit increase.
Updating scaling for data misfits by 2.0334092788677185
New scales: [0.06755054 0.93244946]
13 1.21e+00 2.92e+01 5.84e+01 1.00e+02 8.91e+01 0
geophys. misfits: 17.5 (target 30.0 [True]); 25.4 (target 30.0 [True]) | smallness misfit: 850.7 (target: 200.0 [False])
Beta cooling evaluation: progress: [17.5 25.4]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 10.991967582503591
14 1.21e+00 2.49e+01 6.66e+01 1.06e+02 7.66e+01 0
geophys. misfits: 20.2 (target 30.0 [True]); 26.9 (target 30.0 [True]) | smallness misfit: 648.5 (target: 200.0 [False])
Beta cooling evaluation: progress: [20.2 26.9]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 14.277505994873788
15 1.21e+00 2.65e+01 6.88e+01 1.10e+02 8.91e+01 0
geophys. misfits: 24.2 (target 30.0 [True]); 26.7 (target 30.0 [True]) | smallness misfit: 526.0 (target: 200.0 [False])
Beta cooling evaluation: progress: [24.2 26.7]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 16.883758265490002
16 1.21e+00 2.65e+01 7.04e+01 1.12e+02 8.84e+01 0
geophys. misfits: 32.1 (target 30.0 [False]); 26.4 (target 30.0 [True]) | smallness misfit: 451.2 (target: 200.0 [False])
Beta cooling evaluation: progress: [32.1 26.4]; minimum progress targets: [30. 30.]
Decreasing beta to counter data misfit increase.
Updating scaling for data misfits by 1.1346842315974561
New scales: [0.07595748 0.92404252]
17 6.06e-01 2.69e+01 6.94e+01 6.89e+01 8.72e+01 0
geophys. misfits: 15.0 (target 30.0 [True]); 24.9 (target 30.0 [True]) | smallness misfit: 469.4 (target: 200.0 [False])
Beta cooling evaluation: progress: [15. 24.9]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 27.05027338261105
18 6.06e-01 2.42e+01 8.22e+01 7.40e+01 8.44e+01 0
geophys. misfits: 15.8 (target 30.0 [True]); 24.0 (target 30.0 [True]) | smallness misfit: 408.7 (target: 200.0 [False])
Beta cooling evaluation: progress: [15.8 24. ]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 42.58458383482374
19 6.06e-01 2.34e+01 9.55e+01 8.13e+01 8.85e+01 0
geophys. misfits: 15.3 (target 30.0 [True]); 23.6 (target 30.0 [True]) | smallness misfit: 357.1 (target: 200.0 [False])
Beta cooling evaluation: progress: [15.3 23.6]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 68.90892466880732
20 6.06e-01 2.29e+01 1.15e+02 9.24e+01 1.01e+02 0
geophys. misfits: 14.2 (target 30.0 [True]); 23.9 (target 30.0 [True]) | smallness misfit: 299.5 (target: 200.0 [False])
Beta cooling evaluation: progress: [14.2 23.9]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 116.21018250023924
21 6.06e-01 2.32e+01 1.41e+02 1.09e+02 1.13e+02 0
geophys. misfits: 29.6 (target 30.0 [True]); 23.7 (target 30.0 [True]) | smallness misfit: 242.6 (target: 200.0 [False])
Beta cooling evaluation: progress: [29.6 23.7]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 132.43748856940522
22 6.06e-01 2.41e+01 1.43e+02 1.11e+02 1.07e+02 0
geophys. misfits: 40.5 (target 30.0 [False]); 24.6 (target 30.0 [True]) | smallness misfit: 224.0 (target: 200.0 [False])
Beta cooling evaluation: progress: [40.5 24.6]; minimum progress targets: [30. 30.]
Decreasing beta to counter data misfit increase.
Updating scaling for data misfits by 1.2192871602369773
New scales: [0.09109663 0.90890337]
23 3.03e-01 2.61e+01 1.38e+02 6.78e+01 9.65e+01 0
geophys. misfits: 25.6 (target 30.0 [True]); 25.9 (target 30.0 [True]) | smallness misfit: 200.3 (target: 200.0 [False])
Beta cooling evaluation: progress: [25.6 25.9]; minimum progress targets: [32.4 30. ]
Warming alpha_pgi to favor clustering: 154.13176095405552
24 3.03e-01 2.59e+01 1.40e+02 6.83e+01 8.86e+01 0
geophys. misfits: 24.8 (target 30.0 [True]); 25.5 (target 30.0 [True]) | smallness misfit: 201.2 (target: 200.0 [False])
Beta cooling evaluation: progress: [24.8 25.5]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 183.63450451485932
25 3.03e-01 2.55e+01 1.53e+02 7.19e+01 9.17e+01 0
geophys. misfits: 31.8 (target 30.0 [False]); 25.1 (target 30.0 [True]) | smallness misfit: 175.0 (target: 200.0 [True])
Beta cooling evaluation: progress: [31.8 25.1]; minimum progress targets: [30. 30.]
Decreasing beta to counter data misfit increase.
Updating scaling for data misfits by 1.1950169651765288
New scales: [0.1069618 0.8930382]
26 1.52e-01 2.58e+01 1.50e+02 4.85e+01 9.23e+01 0
geophys. misfits: 20.9 (target 30.0 [True]); 24.5 (target 30.0 [True]) | smallness misfit: 194.2 (target: 200.0 [True])
All targets have been reached
Beta cooling evaluation: progress: [20.9 24.5]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 244.12448308547073
------------------------- STOP! -------------------------
1 : |fc-fOld| = 0.0000e+00 <= tolF*(1+|f0|) = 3.0000e+04
0 : |xc-x_last| = 4.5300e-01 <= tolX*(1+|x0|) = 1.0000e-06
0 : |proj(x-g)-x| = 9.2304e+01 <= tolG = 1.0000e-01
0 : |proj(x-g)-x| = 9.2304e+01 <= 1e3*eps = 1.0000e-02
0 : maxIter = 50 <= iter = 27
------------------------- DONE! -------------------------
Running inversion with SimPEG v0.23.1.dev10+gf697d2455
simpeg.InvProblem will set Regularization.reference_model to m0.
simpeg.InvProblem will set Regularization.reference_model to m0.
simpeg.InvProblem will set Regularization.reference_model to m0.
simpeg.InvProblem is setting bfgsH0 to the inverse of the eval2Deriv.
***Done using the default solver Pardiso and no solver_opts.***
Alpha scales: [np.float64(0.00034582715575284253), np.float64(0.0), np.float64(3.505139062262835e-06), np.float64(0.0)]
Calculating the scaling parameter.
Scale Multipliers: [0.10209669 0.89790331]
<class 'simpeg.regularization.pgi.PGIsmallness'>
Initial data misfit scales: [0.10209669 0.89790331]
model has any nan: 0
=============================== Projected GNCG ===============================
# beta phi_d phi_m f |proj(x-g)-x| LS Comment
-----------------------------------------------------------------------------
x0 has any nan: 0
0 1.94e+03 3.00e+05 0.00e+00 3.00e+05 1.41e+02 0
geophys. misfits: 84973.1 (target 30.0 [False]); 63603.2 (target 30.0 [False]) | smallness misfit: 284.1 (target: 200.0 [False])
Beta cooling evaluation: progress: [84973.1 63603.2]; minimum progress targets: [240000. 240000.]
1 1.94e+03 6.58e+04 6.75e-01 6.71e+04 9.11e+01 0
geophys. misfits: 557.3 (target 30.0 [False]); 29.9 (target 30.0 [True]) | smallness misfit: 99.2 (target: 200.0 [True])
Beta cooling evaluation: progress: [557.3 29.9]; minimum progress targets: [67978.5 50882.6]
Updating scaling for data misfits by 1.0030736670972549
New scales: [0.10237838 0.89762162]
2 1.94e+03 8.39e+01 2.57e-01 5.83e+02 6.99e+01 0 Skip BFGS
geophys. misfits: 100.2 (target 30.0 [False]); 28.8 (target 30.0 [True]) | smallness misfit: 63.0 (target: 200.0 [True])
Beta cooling evaluation: progress: [100.2 28.8]; minimum progress targets: [445.9 30. ]
Updating scaling for data misfits by 1.0423893952972354
New scales: [0.106257 0.893743]
3 1.94e+03 3.64e+01 1.82e-01 3.91e+02 5.85e+01 0 Skip BFGS
geophys. misfits: 98.4 (target 30.0 [False]); 27.3 (target 30.0 [True]) | smallness misfit: 44.9 (target: 200.0 [True])
Beta cooling evaluation: progress: [98.4 27.3]; minimum progress targets: [80.1 30. ]
Decreasing beta to counter data misfit decrase plateau.
Updating scaling for data misfits by 1.1005956441398324
New scales: [0.11570918 0.88429082]
4 9.72e+02 3.55e+01 1.23e-01 1.55e+02 1.02e+02 0
geophys. misfits: 33.0 (target 30.0 [False]); 25.1 (target 30.0 [True]) | smallness misfit: 47.7 (target: 200.0 [True])
Beta cooling evaluation: progress: [33. 25.1]; minimum progress targets: [78.7 30. ]
Updating scaling for data misfits by 1.1953298841946123
New scales: [0.13525371 0.86474629]
5 9.72e+02 2.62e+01 1.30e-01 1.52e+02 4.36e+01 0
geophys. misfits: 27.0 (target 30.0 [True]); 25.3 (target 30.0 [True]) | smallness misfit: 48.1 (target: 200.0 [True])
All targets have been reached
Beta cooling evaluation: progress: [27. 25.3]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 1.1487880303993072
------------------------- STOP! -------------------------
1 : |fc-fOld| = 0.0000e+00 <= tolF*(1+|f0|) = 3.0000e+04
0 : |xc-x_last| = 1.0975e-01 <= tolX*(1+|x0|) = 1.0000e-06
0 : |proj(x-g)-x| = 4.3565e+01 <= tolG = 1.0000e-01
0 : |proj(x-g)-x| = 4.3565e+01 <= 1e3*eps = 1.0000e-02
0 : maxIter = 50 <= iter = 6
------------------------- DONE! -------------------------
Running inversion with SimPEG v0.23.1.dev10+gf697d2455
simpeg.InvProblem will set Regularization.reference_model to m0.
simpeg.InvProblem will set Regularization.reference_model to m0.
simpeg.InvProblem is setting bfgsH0 to the inverse of the eval2Deriv.
***Done using the default solver Pardiso and no solver_opts.***
Alpha scales: [np.float64(3.490296034007822e-05), np.float64(0.0), np.float64(4.7063306913004674e-05), np.float64(0.0)]
Calculating the scaling parameter.
Scale Multipliers: [0.10209669 0.89790331]
/home/vsts/work/1/s/simpeg/directives/directives.py:339: UserWarning:
There is no PGI regularization. Smallness target is turned off (TriggerSmall flag)
Initial data misfit scales: [0.10209669 0.89790331]
model has any nan: 0
=============================== Projected GNCG ===============================
# beta phi_d phi_m f |proj(x-g)-x| LS Comment
-----------------------------------------------------------------------------
x0 has any nan: 0
0 1.04e+06 3.00e+05 0.00e+00 3.00e+05 1.41e+02 0
geophys. misfits: 53322.7 (target 30.0 [False]); 36451.0 (target 30.0 [False])
1 2.08e+05 3.82e+04 4.30e-02 4.71e+04 1.38e+02 0
geophys. misfits: 7580.5 (target 30.0 [False]); 3977.3 (target 30.0 [False])
2 4.16e+04 4.35e+03 1.08e-01 8.82e+03 1.32e+02 0 Skip BFGS
geophys. misfits: 486.8 (target 30.0 [False]); 280.4 (target 30.0 [False])
3 8.31e+03 3.01e+02 1.42e-01 1.48e+03 1.04e+02 0 Skip BFGS
geophys. misfits: 30.9 (target 30.0 [False]); 47.7 (target 30.0 [False])
4 1.66e+03 4.60e+01 1.53e-01 3.00e+02 9.08e+01 0 Skip BFGS
geophys. misfits: 10.6 (target 30.0 [True]); 22.7 (target 30.0 [True])
All targets have been reached
------------------------- STOP! -------------------------
1 : |fc-fOld| = 0.0000e+00 <= tolF*(1+|f0|) = 3.0000e+04
0 : |xc-x_last| = 8.1902e-01 <= tolX*(1+|x0|) = 1.0000e-06
0 : |proj(x-g)-x| = 9.0707e+01 <= tolG = 1.0000e-01
0 : |proj(x-g)-x| = 9.0707e+01 <= 1e3*eps = 1.0000e-02
0 : maxIter = 50 <= iter = 5
------------------------- DONE! -------------------------
/home/vsts/work/1/s/examples/10-pgi/plot_inv_1_PGI_Linear_1D_joint_WithRelationships.py:302: UserWarning:
marker is redundantly defined by the 'marker' keyword argument and the fmt string "b.-" (-> marker='.'). The keyword argument will take precedence.
/home/vsts/work/1/s/examples/10-pgi/plot_inv_1_PGI_Linear_1D_joint_WithRelationships.py:309: UserWarning:
marker is redundantly defined by the 'marker' keyword argument and the fmt string "r.-" (-> marker='.'). The keyword argument will take precedence.
/home/vsts/work/1/s/examples/10-pgi/plot_inv_1_PGI_Linear_1D_joint_WithRelationships.py:353: UserWarning:
marker is redundantly defined by the 'marker' keyword argument and the fmt string "b.-" (-> marker='.'). The keyword argument will take precedence.
/home/vsts/work/1/s/examples/10-pgi/plot_inv_1_PGI_Linear_1D_joint_WithRelationships.py:360: UserWarning:
marker is redundantly defined by the 'marker' keyword argument and the fmt string "r.-" (-> marker='.'). The keyword argument will take precedence.
/home/vsts/work/1/s/examples/10-pgi/plot_inv_1_PGI_Linear_1D_joint_WithRelationships.py:367: UserWarning:
The following kwargs were not used by contour: 'label'
/home/vsts/work/1/s/examples/10-pgi/plot_inv_1_PGI_Linear_1D_joint_WithRelationships.py:412: UserWarning:
marker is redundantly defined by the 'marker' keyword argument and the fmt string "b.-" (-> marker='.'). The keyword argument will take precedence.
/home/vsts/work/1/s/examples/10-pgi/plot_inv_1_PGI_Linear_1D_joint_WithRelationships.py:419: UserWarning:
marker is redundantly defined by the 'marker' keyword argument and the fmt string "r.-" (-> marker='.'). The keyword argument will take precedence.
import discretize as Mesh
import matplotlib.pyplot as plt
import matplotlib.lines as mlines
import numpy as np
from simpeg import (
data_misfit,
directives,
inverse_problem,
inversion,
maps,
optimization,
regularization,
simulation,
utils,
)
# Random seed for reproductibility
np.random.seed(1)
# Mesh
N = 100
mesh = Mesh.TensorMesh([N])
# Survey design parameters
nk = 30
jk = np.linspace(1.0, 59.0, nk)
p = -0.25
q = 0.25
# Physics
def g(k):
return np.exp(p * jk[k] * mesh.cell_centers_x) * np.cos(
np.pi * q * jk[k] * mesh.cell_centers_x
)
G = np.empty((nk, mesh.nC))
for i in range(nk):
G[i, :] = g(i)
m0 = np.zeros(mesh.nC)
m0[20:41] = np.linspace(0.0, 1.0, 21)
m0[41:57] = np.linspace(-1, 0.0, 16)
poly0 = maps.PolynomialPetroClusterMap(coeffyx=np.r_[0.0, -4.0, 4.0])
poly1 = maps.PolynomialPetroClusterMap(coeffyx=np.r_[-0.0, 3.0, 6.0, 6.0])
poly0_inverse = maps.PolynomialPetroClusterMap(coeffyx=-np.r_[0.0, -4.0, 4.0])
poly1_inverse = maps.PolynomialPetroClusterMap(coeffyx=-np.r_[0.0, 3.0, 6.0, 6.0])
cluster_mapping = [maps.IdentityMap(), poly0_inverse, poly1_inverse]
m1 = np.zeros(100)
m1[20:41] = 1.0 + (poly0 * np.vstack([m0[20:41], m1[20:41]]).T)[:, 1]
m1[41:57] = -1.0 + (poly1 * np.vstack([m0[41:57], m1[41:57]]).T)[:, 1]
model2d = np.vstack([m0, m1]).T
m = utils.mkvc(model2d)
clfmapping = utils.GaussianMixtureWithNonlinearRelationships(
mesh=mesh,
n_components=3,
covariance_type="full",
tol=1e-8,
reg_covar=1e-3,
max_iter=1000,
n_init=100,
init_params="kmeans",
random_state=None,
warm_start=False,
means_init=np.array(
[
[0, 0],
[m0[20:41].mean(), m1[20:41].mean()],
[m0[41:57].mean(), m1[41:57].mean()],
]
),
verbose=0,
verbose_interval=10,
cluster_mapping=cluster_mapping,
)
clfmapping = clfmapping.fit(model2d)
clfnomapping = utils.WeightedGaussianMixture(
mesh=mesh,
n_components=3,
covariance_type="full",
tol=1e-8,
reg_covar=1e-3,
max_iter=1000,
n_init=100,
init_params="kmeans",
random_state=None,
warm_start=False,
verbose=0,
verbose_interval=10,
)
clfnomapping = clfnomapping.fit(model2d)
wires = maps.Wires(("m1", mesh.nC), ("m2", mesh.nC))
relatrive_error = 0.01
noise_floor = 0.0
prob1 = simulation.LinearSimulation(mesh, G=G, model_map=wires.m1)
survey1 = prob1.make_synthetic_data(
m, relative_error=relatrive_error, noise_floor=noise_floor, add_noise=True
)
prob2 = simulation.LinearSimulation(mesh, G=G, model_map=wires.m2)
survey2 = prob2.make_synthetic_data(
m, relative_error=relatrive_error, noise_floor=noise_floor, add_noise=True
)
dmis1 = data_misfit.L2DataMisfit(simulation=prob1, data=survey1)
dmis2 = data_misfit.L2DataMisfit(simulation=prob2, data=survey2)
dmis = dmis1 + dmis2
minit = np.zeros_like(m)
# Distance weighting
wr1 = np.sum(prob1.G**2.0, axis=0) ** 0.5 / mesh.cell_volumes
wr1 = wr1 / np.max(wr1)
wr2 = np.sum(prob2.G**2.0, axis=0) ** 0.5 / mesh.cell_volumes
wr2 = wr2 / np.max(wr2)
reg_simple = regularization.PGI(
mesh=mesh,
gmmref=clfmapping,
gmm=clfmapping,
approx_gradient=True,
wiresmap=wires,
non_linear_relationships=True,
weights_list=[wr1, wr2],
)
opt = optimization.ProjectedGNCG(
maxIter=50,
tolX=1e-6,
maxIterCG=100,
tolCG=1e-3,
lower=-10,
upper=10,
)
invProb = inverse_problem.BaseInvProblem(dmis, reg_simple, opt)
# directives
scales = directives.ScalingMultipleDataMisfits_ByEig(
chi0_ratio=np.r_[1.0, 1.0], verbose=True, n_pw_iter=10
)
scaling_schedule = directives.JointScalingSchedule(verbose=True)
alpha0_ratio = np.r_[1e6, 1e4, 1, 1]
alphas = directives.AlphasSmoothEstimate_ByEig(
alpha0_ratio=alpha0_ratio, n_pw_iter=10, verbose=True
)
beta = directives.BetaEstimate_ByEig(beta0_ratio=1e-5, n_pw_iter=10)
betaIt = directives.PGI_BetaAlphaSchedule(
verbose=True,
coolingFactor=2.0,
progress=0.2,
)
targets = directives.MultiTargetMisfits(verbose=True)
petrodir = directives.PGI_UpdateParameters(update_gmm=False)
# Setup Inversion
inv = inversion.BaseInversion(
invProb,
directiveList=[alphas, scales, beta, petrodir, targets, betaIt, scaling_schedule],
)
mcluster_map = inv.run(minit)
# Inversion with no nonlinear mapping
reg_simple_no_map = regularization.PGI(
mesh=mesh,
gmmref=clfnomapping,
gmm=clfnomapping,
approx_gradient=True,
wiresmap=wires,
non_linear_relationships=False,
weights_list=[wr1, wr2],
)
opt = optimization.ProjectedGNCG(
maxIter=50,
tolX=1e-6,
maxIterCG=100,
tolCG=1e-3,
lower=-10,
upper=10,
)
invProb = inverse_problem.BaseInvProblem(dmis, reg_simple_no_map, opt)
# directives
scales = directives.ScalingMultipleDataMisfits_ByEig(
chi0_ratio=np.r_[1.0, 1.0], verbose=True, n_pw_iter=10
)
scaling_schedule = directives.JointScalingSchedule(verbose=True)
alpha0_ratio = np.r_[100.0 * np.ones(2), 1, 1]
alphas = directives.AlphasSmoothEstimate_ByEig(
alpha0_ratio=alpha0_ratio, n_pw_iter=10, verbose=True
)
beta = directives.BetaEstimate_ByEig(beta0_ratio=1e-5, n_pw_iter=10)
betaIt = directives.PGI_BetaAlphaSchedule(
verbose=True,
coolingFactor=2.0,
progress=0.2,
)
targets = directives.MultiTargetMisfits(
chiSmall=1.0, TriggerSmall=True, TriggerTheta=False, verbose=True
)
petrodir = directives.PGI_UpdateParameters(update_gmm=False)
# Setup Inversion
inv = inversion.BaseInversion(
invProb,
directiveList=[alphas, scales, beta, petrodir, targets, betaIt, scaling_schedule],
)
mcluster_no_map = inv.run(minit)
# WeightedLeastSquares Inversion
reg1 = regularization.WeightedLeastSquares(
mesh, alpha_s=1.0, alpha_x=1.0, mapping=wires.m1, weights={"cell_weights": wr1}
)
reg2 = regularization.WeightedLeastSquares(
mesh, alpha_s=1.0, alpha_x=1.0, mapping=wires.m2, weights={"cell_weights": wr2}
)
reg = reg1 + reg2
opt = optimization.ProjectedGNCG(
maxIter=50,
tolX=1e-6,
maxIterCG=100,
tolCG=1e-3,
lower=-10,
upper=10,
)
invProb = inverse_problem.BaseInvProblem(dmis, reg, opt)
# directives
alpha0_ratio = np.r_[1, 1, 1, 1]
alphas = directives.AlphasSmoothEstimate_ByEig(
alpha0_ratio=alpha0_ratio, n_pw_iter=10, verbose=True
)
scales = directives.ScalingMultipleDataMisfits_ByEig(
chi0_ratio=np.r_[1.0, 1.0], verbose=True, n_pw_iter=10
)
scaling_schedule = directives.JointScalingSchedule(verbose=True)
beta = directives.BetaEstimate_ByEig(beta0_ratio=1e-5, n_pw_iter=10)
beta_schedule = directives.BetaSchedule(coolingFactor=5.0, coolingRate=1)
targets = directives.MultiTargetMisfits(
TriggerSmall=False,
verbose=True,
)
# Setup Inversion
inv = inversion.BaseInversion(
invProb,
directiveList=[alphas, scales, beta, targets, beta_schedule, scaling_schedule],
)
mtik = inv.run(minit)
# Final Plot
fig, axes = plt.subplots(3, 4, figsize=(25, 15))
axes = axes.reshape(12)
left, width = 0.25, 0.5
bottom, height = 0.25, 0.5
right = left + width
top = bottom + height
axes[0].set_axis_off()
axes[0].text(
0.5 * (left + right),
0.5 * (bottom + top),
("Using true nonlinear\npetrophysical relationships"),
horizontalalignment="center",
verticalalignment="center",
fontsize=20,
color="black",
transform=axes[0].transAxes,
)
axes[1].plot(mesh.cell_centers_x, wires.m1 * mcluster_map, "b.-", ms=5, marker="v")
axes[1].plot(mesh.cell_centers_x, wires.m1 * m, "k--")
axes[1].set_title("Problem 1")
axes[1].legend(["Recovered Model", "True Model"], loc=1)
axes[1].set_xlabel("X")
axes[1].set_ylabel("Property 1")
axes[2].plot(mesh.cell_centers_x, wires.m2 * mcluster_map, "r.-", ms=5, marker="v")
axes[2].plot(mesh.cell_centers_x, wires.m2 * m, "k--")
axes[2].set_title("Problem 2")
axes[2].legend(["Recovered Model", "True Model"], loc=1)
axes[2].set_xlabel("X")
axes[2].set_ylabel("Property 2")
x, y = np.mgrid[-1:1:0.01, -4:2:0.01]
pos = np.empty(x.shape + (2,))
pos[:, :, 0] = x
pos[:, :, 1] = y
CS = axes[3].contour(
x,
y,
np.exp(clfmapping.score_samples(pos.reshape(-1, 2)).reshape(x.shape)),
100,
alpha=0.25,
cmap="viridis",
)
cs_proxy = mlines.Line2D([], [], label="True Petrophysical Distribution")
ps = axes[3].scatter(
wires.m1 * mcluster_map,
wires.m2 * mcluster_map,
marker="v",
label="Recovered model crossplot",
)
axes[3].set_title("Petrophysical Distribution")
axes[3].legend(handles=[cs_proxy, ps])
axes[3].set_xlabel("Property 1")
axes[3].set_ylabel("Property 2")
axes[4].set_axis_off()
axes[4].text(
0.5 * (left + right),
0.5 * (bottom + top),
("Using a pure\nGaussian distribution"),
horizontalalignment="center",
verticalalignment="center",
fontsize=20,
color="black",
transform=axes[4].transAxes,
)
axes[5].plot(mesh.cell_centers_x, wires.m1 * mcluster_no_map, "b.-", ms=5, marker="v")
axes[5].plot(mesh.cell_centers_x, wires.m1 * m, "k--")
axes[5].set_title("Problem 1")
axes[5].legend(["Recovered Model", "True Model"], loc=1)
axes[5].set_xlabel("X")
axes[5].set_ylabel("Property 1")
axes[6].plot(mesh.cell_centers_x, wires.m2 * mcluster_no_map, "r.-", ms=5, marker="v")
axes[6].plot(mesh.cell_centers_x, wires.m2 * m, "k--")
axes[6].set_title("Problem 2")
axes[6].legend(["Recovered Model", "True Model"], loc=1)
axes[6].set_xlabel("X")
axes[6].set_ylabel("Property 2")
CSF = axes[7].contour(
x,
y,
np.exp(clfmapping.score_samples(pos.reshape(-1, 2)).reshape(x.shape)),
100,
alpha=0.5,
label="True Petro. Distribution",
)
CS = axes[7].contour(
x,
y,
np.exp(clfnomapping.score_samples(pos.reshape(-1, 2)).reshape(x.shape)),
500,
cmap="viridis",
linestyles="--",
)
axes[7].scatter(
wires.m1 * mcluster_no_map,
wires.m2 * mcluster_no_map,
marker="v",
label="Recovered model crossplot",
)
cs_modeled_proxy = mlines.Line2D(
[], [], linestyle="--", label="Modeled Petro. Distribution"
)
axes[7].set_title("Petrophysical Distribution")
axes[7].legend(handles=[cs_proxy, cs_modeled_proxy, ps])
axes[7].set_xlabel("Property 1")
axes[7].set_ylabel("Property 2")
# Tikonov
axes[8].set_axis_off()
axes[8].text(
0.5 * (left + right),
0.5 * (bottom + top),
("Least-Squares\n~Using a single cluster"),
horizontalalignment="center",
verticalalignment="center",
fontsize=20,
color="black",
transform=axes[8].transAxes,
)
axes[9].plot(mesh.cell_centers_x, wires.m1 * mtik, "b.-", ms=5, marker="v")
axes[9].plot(mesh.cell_centers_x, wires.m1 * m, "k--")
axes[9].set_title("Problem 1")
axes[9].legend(["Recovered Model", "True Model"], loc=1)
axes[9].set_xlabel("X")
axes[9].set_ylabel("Property 1")
axes[10].plot(mesh.cell_centers_x, wires.m2 * mtik, "r.-", ms=5, marker="v")
axes[10].plot(mesh.cell_centers_x, wires.m2 * m, "k--")
axes[10].set_title("Problem 2")
axes[10].legend(["Recovered Model", "True Model"], loc=1)
axes[10].set_xlabel("X")
axes[10].set_ylabel("Property 2")
CS = axes[11].contour(
x,
y,
np.exp(clfmapping.score_samples(pos.reshape(-1, 2)).reshape(x.shape)),
100,
alpha=0.25,
cmap="viridis",
)
axes[11].scatter(wires.m1 * mtik, wires.m2 * mtik, marker="v")
axes[11].set_title("Petro Distribution")
axes[11].legend(handles=[cs_proxy, ps])
axes[11].set_xlabel("Property 1")
axes[11].set_ylabel("Property 2")
plt.subplots_adjust(wspace=0.3, hspace=0.3, top=0.85)
plt.show()
Total running time of the script: (0 minutes 56.493 seconds)
Estimated memory usage: 293 MB
