Note
Go to the end to download the full example code.
Petrophysically guided inversion: Joint linear example with nonlinear relationships#
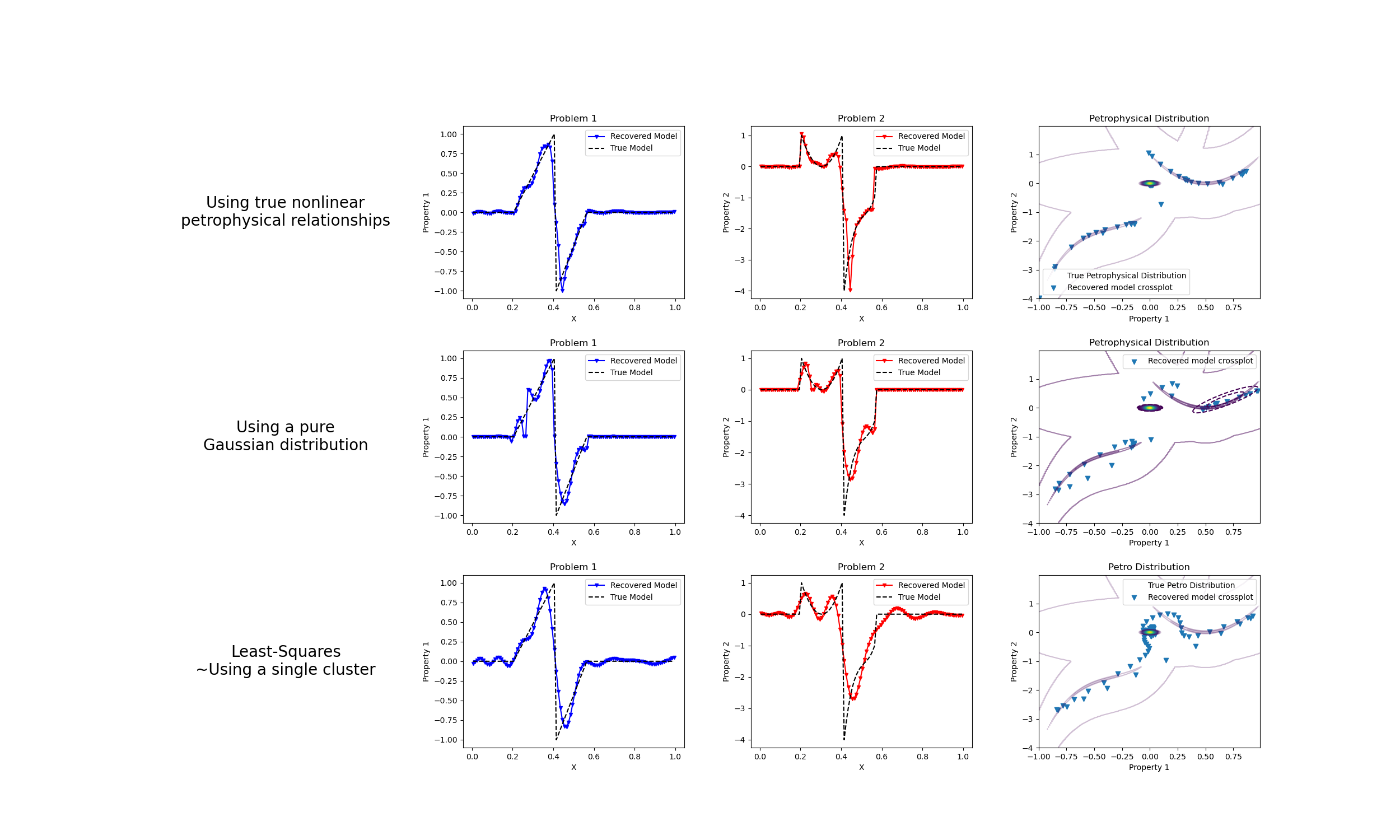
We do a comparison between the classic least-squares inversion and our formulation of a petrophysically guided inversion. We explore it through coupling two linear problems whose respective physical properties are linked by polynomial relationships that change between rock units.
Running inversion with SimPEG v0.25.2
Alpha scales: [np.float64(3.479046834083167), np.float64(0.0), np.float64(3.476000760552e-06), np.float64(0.0)]
Calculating the scaling parameter.
Scale Multipliers: [0.09589118 0.90410882]
<class 'simpeg.regularization.pgi.PGIsmallness'>
Initial data misfit scales: [0.09589118 0.90410882]
================================================= Projected GNCG =================================================
# beta phi_d phi_m f |proj(x-g)-x| LS iter_CG CG |Ax-b|/|b| CG |Ax-b| Comment
-----------------------------------------------------------------------------------------------------------------
0 1.87e+01 3.00e+05 0.00e+00 3.00e+05 0 inf inf
1 1.87e+01 1.46e+03 1.71e+02 4.65e+03 1.41e+02 0 20 8.28e-04 7.54e+03
geophys. misfits: 8154.3 (target 30.0 [False]); 752.7 (target 30.0 [False]) | smallness misfit: 4038.3 (target: 200.0 [False])
Beta cooling evaluation: progress: [8154.3 752.7]; minimum progress targets: [240000. 240000.]
2 1.87e+01 6.30e+01 4.09e+01 8.28e+02 1.39e+02 0 100 3.59e-03 2.70e+01 Skip BFGS
geophys. misfits: 472.1 (target 30.0 [False]); 19.6 (target 30.0 [True]) | smallness misfit: 1393.4 (target: 200.0 [False])
Beta cooling evaluation: progress: [472.1 19.6]; minimum progress targets: [6523.4 602.1]
Updating scaling for data misfits by 1.5307801410415616
New scales: [0.13967904 0.86032096]
3 1.87e+01 5.59e+01 4.12e+01 8.26e+02 7.92e+01 0 100 3.57e+00 9.98e+02 Skip BFGS
geophys. misfits: 283.1 (target 30.0 [False]); 19.1 (target 30.0 [True]) | smallness misfit: 1162.4 (target: 200.0 [False])
Beta cooling evaluation: progress: [283.1 19.1]; minimum progress targets: [377.7 30. ]
Updating scaling for data misfits by 1.573493423851348
New scales: [0.20348397 0.79651603]
4 1.87e+01 4.93e+01 4.22e+01 8.38e+02 9.42e+01 0 100 1.16e-02 1.12e+01
geophys. misfits: 167.8 (target 30.0 [False]); 19.0 (target 30.0 [True]) | smallness misfit: 1111.4 (target: 200.0 [False])
Beta cooling evaluation: progress: [167.8 19. ]; minimum progress targets: [226.5 30. ]
Updating scaling for data misfits by 1.5808988700350344
New scales: [0.28768247 0.71231753]
5 1.87e+01 4.41e+01 4.30e+01 8.48e+02 7.49e+01 0 100 2.06e-01 5.69e+01 Skip BFGS
geophys. misfits: 105.5 (target 30.0 [False]); 19.3 (target 30.0 [True]) | smallness misfit: 1067.7 (target: 200.0 [False])
Beta cooling evaluation: progress: [105.5 19.3]; minimum progress targets: [134.2 30. ]
Updating scaling for data misfits by 1.5542048869507739
New scales: [0.38563395 0.61436605]
6 1.87e+01 4.04e+01 4.35e+01 8.55e+02 7.79e+01 0 100 3.14e-01 7.71e+01 Skip BFGS
geophys. misfits: 73.0 (target 30.0 [False]); 20.0 (target 30.0 [True]) | smallness misfit: 1033.2 (target: 200.0 [False])
Beta cooling evaluation: progress: [73. 20.]; minimum progress targets: [84.4 30. ]
Updating scaling for data misfits by 1.5026400280403849
New scales: [0.48538448 0.51461552]
7 1.87e+01 3.80e+01 4.39e+01 8.59e+02 7.92e+01 0 100 1.83e+00 3.69e+02 Skip BFGS
geophys. misfits: 56.0 (target 30.0 [False]); 21.1 (target 30.0 [True]) | smallness misfit: 1002.6 (target: 200.0 [False])
Beta cooling evaluation: progress: [56. 21.1]; minimum progress targets: [58.4 30. ]
Updating scaling for data misfits by 1.4250778031167355
New scales: [0.57340267 0.42659733]
8 1.87e+01 3.65e+01 4.41e+01 8.61e+02 8.30e+01 0 100 4.48e-01 1.51e+02 Skip BFGS
geophys. misfits: 46.8 (target 30.0 [False]); 22.5 (target 30.0 [True]) | smallness misfit: 975.5 (target: 200.0 [False])
Beta cooling evaluation: progress: [46.8 22.5]; minimum progress targets: [44.8 30. ]
Decreasing beta to counter data misfit decrase plateau.
Updating scaling for data misfits by 1.331701051431981
New scales: [0.64157457 0.35842543]
9 9.35e+00 2.20e+01 4.52e+01 4.45e+02 9.26e+01 0 100 1.06e-01 5.08e+01
geophys. misfits: 24.1 (target 30.0 [True]); 18.4 (target 30.0 [True]) | smallness misfit: 1015.4 (target: 200.0 [False])
Beta cooling evaluation: progress: [24.1 18.4]; minimum progress targets: [37.5 30. ]
Warming alpha_pgi to favor clustering: 1.4400341944398984
10 9.35e+00 2.26e+01 4.61e+01 4.54e+02 5.02e+01 0 100 7.69e-01 5.11e+01
geophys. misfits: 23.9 (target 30.0 [True]); 20.4 (target 30.0 [True]) | smallness misfit: 936.7 (target: 200.0 [False])
Beta cooling evaluation: progress: [23.9 20.4]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 1.9645442724486077
11 9.35e+00 2.35e+01 4.71e+01 4.64e+02 5.26e+01 0 100 3.31e-01 2.30e+01
geophys. misfits: 23.9 (target 30.0 [True]); 22.8 (target 30.0 [True]) | smallness misfit: 873.0 (target: 200.0 [False])
Beta cooling evaluation: progress: [23.9 22.8]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 2.5247257560938823
12 9.35e+00 2.45e+01 4.81e+01 4.74e+02 4.11e+01 0 100 7.38e+00 3.98e+02 Skip BFGS
geophys. misfits: 23.9 (target 30.0 [True]); 25.5 (target 30.0 [True]) | smallness misfit: 818.1 (target: 200.0 [False])
Beta cooling evaluation: progress: [23.9 25.5]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 3.071816100389489
13 9.35e+00 2.53e+01 4.90e+01 4.84e+02 6.39e+01 0 100 9.66e-01 3.88e+02 Skip BFGS
geophys. misfits: 23.8 (target 30.0 [True]); 28.1 (target 30.0 [True]) | smallness misfit: 774.0 (target: 200.0 [False])
Beta cooling evaluation: progress: [23.8 28.1]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 3.5755791048725625
14 9.35e+00 2.62e+01 4.98e+01 4.92e+02 8.07e+01 0 100 1.22e-01 4.78e+01 Skip BFGS
geophys. misfits: 23.8 (target 30.0 [True]); 30.6 (target 30.0 [False]) | smallness misfit: 738.7 (target: 200.0 [False])
Beta cooling evaluation: progress: [23.8 30.6]; minimum progress targets: [30. 30.]
Decreasing beta to counter data misfit increase.
Updating scaling for data misfits by 1.2622428610466754
New scales: [0.58645136 0.41354864]
15 4.68e+00 1.89e+01 5.09e+01 2.57e+02 9.29e+01 0 100 1.27e+01 2.80e+03
geophys. misfits: 18.3 (target 30.0 [True]); 19.7 (target 30.0 [True]) | smallness misfit: 819.6 (target: 200.0 [False])
Beta cooling evaluation: progress: [18.3 19.7]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 5.660880794436028
16 4.68e+00 2.03e+01 5.42e+01 2.74e+02 8.33e+01 0 100 6.81e-02 1.91e+02
geophys. misfits: 18.4 (target 30.0 [True]); 22.9 (target 30.0 [True]) | smallness misfit: 706.4 (target: 200.0 [False])
Beta cooling evaluation: progress: [18.4 22.9]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 8.324260166957588
17 4.68e+00 2.20e+01 5.80e+01 2.93e+02 7.87e+01 0 100 1.33e+00 3.11e+02
geophys. misfits: 18.8 (target 30.0 [True]); 26.5 (target 30.0 [True]) | smallness misfit: 626.6 (target: 200.0 [False])
Beta cooling evaluation: progress: [18.8 26.5]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 11.348125663370787
18 4.68e+00 2.38e+01 6.19e+01 3.13e+02 9.54e+01 0 100 1.96e+00 6.80e+02
geophys. misfits: 19.3 (target 30.0 [True]); 30.2 (target 30.0 [False]) | smallness misfit: 560.1 (target: 200.0 [False])
Beta cooling evaluation: progress: [19.3 30.2]; minimum progress targets: [30. 30.]
Decreasing beta to counter data misfit increase.
Updating scaling for data misfits by 1.5545841115723387
New scales: [0.47704278 0.52295722]
19 2.34e+00 1.81e+01 6.37e+01 1.67e+02 9.81e+01 0 100 2.13e+00 1.76e+03
geophys. misfits: 17.0 (target 30.0 [True]); 19.0 (target 30.0 [True]) | smallness misfit: 608.2 (target: 200.0 [False])
Beta cooling evaluation: progress: [17. 19.]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 18.942662046986733
20 2.34e+00 2.03e+01 7.27e+01 1.90e+02 9.93e+01 0 100 2.27e+00 4.03e+03
geophys. misfits: 17.7 (target 30.0 [True]); 22.7 (target 30.0 [True]) | smallness misfit: 494.2 (target: 200.0 [False])
Beta cooling evaluation: progress: [17.7 22.7]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 28.528342635446194
21 2.34e+00 1.94e+01 8.31e+01 2.14e+02 1.03e+02 0 100 5.15e-01 2.08e+03
geophys. misfits: 17.8 (target 30.0 [True]); 21.0 (target 30.0 [True]) | smallness misfit: 419.6 (target: 200.0 [False])
Beta cooling evaluation: progress: [17.8 21. ]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 44.483092128227675
22 2.34e+00 2.07e+01 9.68e+01 2.47e+02 1.08e+02 0 100 5.40e+00 1.14e+04
geophys. misfits: 18.8 (target 30.0 [True]); 22.4 (target 30.0 [True]) | smallness misfit: 337.4 (target: 200.0 [False])
Beta cooling evaluation: progress: [18.8 22.4]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 65.34984123149157
23 2.34e+00 2.67e+01 1.09e+02 2.82e+02 1.19e+02 0 100 1.26e+00 1.44e+04
geophys. misfits: 19.6 (target 30.0 [True]); 33.1 (target 30.0 [False]) | smallness misfit: 309.5 (target: 200.0 [False])
Beta cooling evaluation: progress: [19.6 33.1]; minimum progress targets: [30. 30.]
Decreasing beta to counter data misfit increase.
Updating scaling for data misfits by 1.53235469423943
New scales: [0.37315649 0.62684351]
24 1.17e+00 2.30e+01 1.10e+02 1.51e+02 1.17e+02 0 100 2.84e-01 3.92e+03
geophys. misfits: 21.5 (target 30.0 [True]); 24.0 (target 30.0 [True]) | smallness misfit: 280.4 (target: 200.0 [False])
Beta cooling evaluation: progress: [21.5 24. ]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 86.56283187335441
25 1.17e+00 2.04e+01 1.24e+02 1.65e+02 9.85e+01 0 100 1.50e+00 5.87e+03
geophys. misfits: 20.7 (target 30.0 [True]); 20.3 (target 30.0 [True]) | smallness misfit: 274.5 (target: 200.0 [False])
Beta cooling evaluation: progress: [20.7 20.3]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 126.71619069258912
26 1.17e+00 2.02e+01 1.49e+02 1.94e+02 1.07e+02 0 100 7.17e-01 4.22e+03
geophys. misfits: 21.1 (target 30.0 [True]); 19.6 (target 30.0 [True]) | smallness misfit: 249.7 (target: 200.0 [False])
Beta cooling evaluation: progress: [21.1 19.6]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 186.82072765944434
27 1.17e+00 2.36e+01 1.80e+02 2.34e+02 1.14e+02 0 100 1.61e+00 6.88e+03
geophys. misfits: 27.5 (target 30.0 [True]); 21.3 (target 30.0 [True]) | smallness misfit: 225.8 (target: 200.0 [False])
Beta cooling evaluation: progress: [27.5 21.3]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 233.335464769764
28 1.17e+00 2.64e+01 1.96e+02 2.55e+02 1.29e+02 0 100 1.82e+00 1.26e+04
geophys. misfits: 31.6 (target 30.0 [False]); 23.4 (target 30.0 [True]) | smallness misfit: 191.5 (target: 200.0 [True])
Beta cooling evaluation: progress: [31.6 23.4]; minimum progress targets: [30. 30.]
Decreasing beta to counter data misfit increase.
Updating scaling for data misfits by 1.282412448684976
New scales: [0.43291789 0.56708211]
29 5.85e-01 2.02e+01 2.04e+02 1.40e+02 1.14e+02 0 100 6.19e-01 7.98e+03
geophys. misfits: 21.9 (target 30.0 [True]); 18.9 (target 30.0 [True]) | smallness misfit: 209.8 (target: 200.0 [False])
Beta cooling evaluation: progress: [21.9 18.9]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 344.931906301962
30 5.85e-01 2.02e+01 2.49e+02 1.66e+02 1.18e+02 1 100 9.37e-01 7.49e+03
geophys. misfits: 21.2 (target 30.0 [True]); 19.4 (target 30.0 [True]) | smallness misfit: 188.1 (target: 200.0 [True])
All targets have been reached
Beta cooling evaluation: progress: [21.2 19.4]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 510.5775688753135
------------------------- STOP! -------------------------
1 : |fc-fOld| = 7.7603e+00 <= tolF*(1+|f0|) = 3.0000e+04
0 : |xc-x_last| = 6.5444e-01 <= tolX*(1+|x0|) = 1.0000e-06
0 : |proj(x-g)-x| = 1.1820e+02 <= tolG = 1.0000e-01
0 : |proj(x-g)-x| = 1.1820e+02 <= 1e3*eps = 1.0000e-02
0 : maxIter = 50 <= iter = 30
------------------------- DONE! -------------------------
Running inversion with SimPEG v0.25.2
Alpha scales: [np.float64(0.0003495886963340754), np.float64(0.0), np.float64(3.4981920061121644e-06), np.float64(0.0)]
Calculating the scaling parameter.
Scale Multipliers: [0.09589118 0.90410882]
<class 'simpeg.regularization.pgi.PGIsmallness'>
Initial data misfit scales: [0.09589118 0.90410882]
================================================= Projected GNCG =================================================
# beta phi_d phi_m f |proj(x-g)-x| LS iter_CG CG |Ax-b|/|b| CG |Ax-b| Comment
-----------------------------------------------------------------------------------------------------------------
0 1.88e+03 3.00e+05 0.00e+00 3.00e+05 0 inf inf
1 1.88e+03 6.44e+04 2.33e+01 1.08e+05 1.41e+02 0 15 3.60e-04 3.27e+03
geophys. misfits: 91168.0 (target 30.0 [False]); 61600.8 (target 30.0 [False]) | smallness misfit: 255.7 (target: 200.0 [False])
Beta cooling evaluation: progress: [91168. 61600.8]; minimum progress targets: [240000. 240000.]
2 1.88e+03 7.20e+01 5.55e-01 1.12e+03 1.35e+02 0 100 8.78e-03 2.35e+02 Skip BFGS
geophys. misfits: 569.7 (target 30.0 [False]); 19.2 (target 30.0 [True]) | smallness misfit: 103.7 (target: 200.0 [True])
Beta cooling evaluation: progress: [569.7 19.2]; minimum progress targets: [72934.4 49280.6]
Updating scaling for data misfits by 1.5596636676672044
New scales: [0.14194049 0.85805951]
3 1.88e+03 2.52e+01 1.36e-01 2.82e+02 1.06e+02 0 100 1.11e-02 2.42e+01 Skip BFGS
geophys. misfits: 75.4 (target 30.0 [False]); 16.9 (target 30.0 [True]) | smallness misfit: 75.3 (target: 200.0 [True])
Beta cooling evaluation: progress: [75.4 16.9]; minimum progress targets: [455.8 30. ]
Updating scaling for data misfits by 1.7804147757468272
New scales: [0.22751099 0.77248901]
4 1.88e+03 2.34e+01 1.32e-01 2.72e+02 8.23e+01 0 100 2.45e-01 3.63e+02
geophys. misfits: 42.4 (target 30.0 [False]); 17.7 (target 30.0 [True]) | smallness misfit: 49.1 (target: 200.0 [True])
Beta cooling evaluation: progress: [42.4 17.7]; minimum progress targets: [60.3 30. ]
Updating scaling for data misfits by 1.6914606856398744
New scales: [0.33251615 0.66748385]
5 1.88e+03 2.26e+01 1.34e-01 2.74e+02 8.72e+01 0 100 1.40e-01 5.01e+01 Skip BFGS
geophys. misfits: 30.4 (target 30.0 [False]); 18.6 (target 30.0 [True]) | smallness misfit: 55.5 (target: 200.0 [True])
Beta cooling evaluation: progress: [30.4 18.6]; minimum progress targets: [34. 30.]
Updating scaling for data misfits by 1.6091335706039764
New scales: [0.44494146 0.55505854]
6 1.88e+03 2.19e+01 1.33e-01 2.72e+02 8.26e+01 0 100 2.14e-02 1.12e+01
geophys. misfits: 24.5 (target 30.0 [True]); 19.8 (target 30.0 [True]) | smallness misfit: 53.3 (target: 200.0 [True])
All targets have been reached
Beta cooling evaluation: progress: [24.5 19.8]; minimum progress targets: [30. 30.]
Warming alpha_pgi to favor clustering: 1.3675655857493167
------------------------- STOP! -------------------------
1 : |fc-fOld| = 2.0521e+01 <= tolF*(1+|f0|) = 3.0000e+04
0 : |xc-x_last| = 1.0833e-01 <= tolX*(1+|x0|) = 1.0000e-06
0 : |proj(x-g)-x| = 8.2587e+01 <= tolG = 1.0000e-01
0 : |proj(x-g)-x| = 8.2587e+01 <= 1e3*eps = 1.0000e-02
0 : maxIter = 50 <= iter = 6
------------------------- DONE! -------------------------
Running inversion with SimPEG v0.25.2
Alpha scales: [np.float64(3.435655378153807e-05), np.float64(0.0), np.float64(3.434575837437124e-05), np.float64(0.0)]
Calculating the scaling parameter.
Scale Multipliers: [0.09589118 0.90410882]
/home/vsts/work/1/s/simpeg/directives/_directives.py:334: UserWarning: There is no PGI regularization. Smallness target is turned off (TriggerSmall flag)
getattr(r, ruleType)()
Initial data misfit scales: [0.09589118 0.90410882]
================================================= Projected GNCG =================================================
# beta phi_d phi_m f |proj(x-g)-x| LS iter_CG CG |Ax-b|/|b| CG |Ax-b| Comment
-----------------------------------------------------------------------------------------------------------------
0 1.02e+06 3.00e+05 0.00e+00 3.00e+05 0 inf inf
1 1.02e+06 4.04e+04 4.30e-02 8.44e+04 1.41e+02 0 24 9.40e-04 8.55e+03
geophys. misfits: 59272.5 (target 30.0 [False]); 38384.6 (target 30.0 [False])
2 2.04e+05 4.18e+03 1.08e-01 2.61e+04 1.37e+02 0 100 3.60e-03 6.57e+01 Skip BFGS
geophys. misfits: 8149.5 (target 30.0 [False]); 3760.7 (target 30.0 [False])
3 4.09e+04 2.68e+02 1.41e-01 6.04e+03 1.30e+02 0 100 6.30e-03 3.17e+01 Skip BFGS
geophys. misfits: 541.2 (target 30.0 [False]); 238.5 (target 30.0 [False])
4 8.17e+03 2.88e+01 1.51e-01 1.27e+03 1.03e+02 0 100 6.76e-01 7.77e+02 Skip BFGS
geophys. misfits: 42.4 (target 30.0 [False]); 27.3 (target 30.0 [True])
Updating scaling for data misfits by 1.0978576555054818
New scales: [0.10429618 0.89570382]
5 1.63e+03 1.50e+01 1.54e-01 2.67e+02 8.79e+01 0 100 2.00e+00 1.60e+03 Skip BFGS
geophys. misfits: 16.9 (target 30.0 [True]); 14.8 (target 30.0 [True])
All targets have been reached
------------------------- STOP! -------------------------
1 : |fc-fOld| = 9.0843e+00 <= tolF*(1+|f0|) = 3.0000e+04
0 : |xc-x_last| = 3.0452e-01 <= tolX*(1+|x0|) = 1.0000e-06
0 : |proj(x-g)-x| = 8.7893e+01 <= tolG = 1.0000e-01
0 : |proj(x-g)-x| = 8.7893e+01 <= 1e3*eps = 1.0000e-02
0 : maxIter = 50 <= iter = 5
------------------------- DONE! -------------------------
/home/vsts/work/1/s/examples/10-pgi/plot_inv_1_PGI_Linear_1D_joint_WithRelationships.py:302: UserWarning: marker is redundantly defined by the 'marker' keyword argument and the fmt string "b.-" (-> marker='.'). The keyword argument will take precedence.
axes[1].plot(mesh.cell_centers_x, wires.m1 * mcluster_map, "b.-", ms=5, marker="v")
/home/vsts/work/1/s/examples/10-pgi/plot_inv_1_PGI_Linear_1D_joint_WithRelationships.py:309: UserWarning: marker is redundantly defined by the 'marker' keyword argument and the fmt string "r.-" (-> marker='.'). The keyword argument will take precedence.
axes[2].plot(mesh.cell_centers_x, wires.m2 * mcluster_map, "r.-", ms=5, marker="v")
/home/vsts/work/1/s/examples/10-pgi/plot_inv_1_PGI_Linear_1D_joint_WithRelationships.py:353: UserWarning: marker is redundantly defined by the 'marker' keyword argument and the fmt string "b.-" (-> marker='.'). The keyword argument will take precedence.
axes[5].plot(mesh.cell_centers_x, wires.m1 * mcluster_no_map, "b.-", ms=5, marker="v")
/home/vsts/work/1/s/examples/10-pgi/plot_inv_1_PGI_Linear_1D_joint_WithRelationships.py:360: UserWarning: marker is redundantly defined by the 'marker' keyword argument and the fmt string "r.-" (-> marker='.'). The keyword argument will take precedence.
axes[6].plot(mesh.cell_centers_x, wires.m2 * mcluster_no_map, "r.-", ms=5, marker="v")
/home/vsts/work/1/s/examples/10-pgi/plot_inv_1_PGI_Linear_1D_joint_WithRelationships.py:367: UserWarning: The following kwargs were not used by contour: 'label'
CSF = axes[7].contour(
/home/vsts/work/1/s/examples/10-pgi/plot_inv_1_PGI_Linear_1D_joint_WithRelationships.py:412: UserWarning: marker is redundantly defined by the 'marker' keyword argument and the fmt string "b.-" (-> marker='.'). The keyword argument will take precedence.
axes[9].plot(mesh.cell_centers_x, wires.m1 * mtik, "b.-", ms=5, marker="v")
/home/vsts/work/1/s/examples/10-pgi/plot_inv_1_PGI_Linear_1D_joint_WithRelationships.py:419: UserWarning: marker is redundantly defined by the 'marker' keyword argument and the fmt string "r.-" (-> marker='.'). The keyword argument will take precedence.
axes[10].plot(mesh.cell_centers_x, wires.m2 * mtik, "r.-", ms=5, marker="v")
import discretize as Mesh
import matplotlib.pyplot as plt
import matplotlib.lines as mlines
import numpy as np
from simpeg import (
data_misfit,
directives,
inverse_problem,
inversion,
maps,
optimization,
regularization,
simulation,
utils,
)
# Random seed for reproductibility
np.random.seed(1)
# Mesh
N = 100
mesh = Mesh.TensorMesh([N])
# Survey design parameters
nk = 30
jk = np.linspace(1.0, 59.0, nk)
p = -0.25
q = 0.25
# Physics
def g(k):
return np.exp(p * jk[k] * mesh.cell_centers_x) * np.cos(
np.pi * q * jk[k] * mesh.cell_centers_x
)
G = np.empty((nk, mesh.nC))
for i in range(nk):
G[i, :] = g(i)
m0 = np.zeros(mesh.nC)
m0[20:41] = np.linspace(0.0, 1.0, 21)
m0[41:57] = np.linspace(-1, 0.0, 16)
poly0 = maps.PolynomialPetroClusterMap(coeffyx=np.r_[0.0, -4.0, 4.0])
poly1 = maps.PolynomialPetroClusterMap(coeffyx=np.r_[-0.0, 3.0, 6.0, 6.0])
poly0_inverse = maps.PolynomialPetroClusterMap(coeffyx=-np.r_[0.0, -4.0, 4.0])
poly1_inverse = maps.PolynomialPetroClusterMap(coeffyx=-np.r_[0.0, 3.0, 6.0, 6.0])
cluster_mapping = [maps.IdentityMap(), poly0_inverse, poly1_inverse]
m1 = np.zeros(100)
m1[20:41] = 1.0 + (poly0 * np.vstack([m0[20:41], m1[20:41]]).T)[:, 1]
m1[41:57] = -1.0 + (poly1 * np.vstack([m0[41:57], m1[41:57]]).T)[:, 1]
model2d = np.vstack([m0, m1]).T
m = utils.mkvc(model2d)
clfmapping = utils.GaussianMixtureWithNonlinearRelationships(
mesh=mesh,
n_components=3,
covariance_type="full",
tol=1e-8,
reg_covar=1e-3,
max_iter=1000,
n_init=100,
init_params="kmeans",
random_state=None,
warm_start=False,
means_init=np.array(
[
[0, 0],
[m0[20:41].mean(), m1[20:41].mean()],
[m0[41:57].mean(), m1[41:57].mean()],
]
),
verbose=0,
verbose_interval=10,
cluster_mapping=cluster_mapping,
)
clfmapping = clfmapping.fit(model2d)
clfnomapping = utils.WeightedGaussianMixture(
mesh=mesh,
n_components=3,
covariance_type="full",
tol=1e-8,
reg_covar=1e-3,
max_iter=1000,
n_init=100,
init_params="kmeans",
random_state=None,
warm_start=False,
verbose=0,
verbose_interval=10,
)
clfnomapping = clfnomapping.fit(model2d)
wires = maps.Wires(("m1", mesh.nC), ("m2", mesh.nC))
relatrive_error = 0.01
noise_floor = 0.0
prob1 = simulation.LinearSimulation(mesh, G=G, model_map=wires.m1)
survey1 = prob1.make_synthetic_data(
m, relative_error=relatrive_error, noise_floor=noise_floor, add_noise=True
)
prob2 = simulation.LinearSimulation(mesh, G=G, model_map=wires.m2)
survey2 = prob2.make_synthetic_data(
m, relative_error=relatrive_error, noise_floor=noise_floor, add_noise=True
)
dmis1 = data_misfit.L2DataMisfit(simulation=prob1, data=survey1)
dmis2 = data_misfit.L2DataMisfit(simulation=prob2, data=survey2)
dmis = dmis1 + dmis2
minit = np.zeros_like(m)
# Distance weighting
wr1 = np.sum(prob1.G**2.0, axis=0) ** 0.5 / mesh.cell_volumes
wr1 = wr1 / np.max(wr1)
wr2 = np.sum(prob2.G**2.0, axis=0) ** 0.5 / mesh.cell_volumes
wr2 = wr2 / np.max(wr2)
reg_simple = regularization.PGI(
mesh=mesh,
gmmref=clfmapping,
gmm=clfmapping,
approx_gradient=True,
wiresmap=wires,
non_linear_relationships=True,
weights_list=[wr1, wr2],
)
opt = optimization.ProjectedGNCG(
maxIter=50,
tolX=1e-6,
cg_maxiter=100,
cg_rtol=1e-3,
lower=-10,
upper=10,
)
invProb = inverse_problem.BaseInvProblem(dmis, reg_simple, opt)
# directives
scales = directives.ScalingMultipleDataMisfits_ByEig(
chi0_ratio=np.r_[1.0, 1.0], verbose=True, n_pw_iter=10
)
scaling_schedule = directives.JointScalingSchedule(verbose=True)
alpha0_ratio = np.r_[1e6, 1e4, 1, 1]
alphas = directives.AlphasSmoothEstimate_ByEig(
alpha0_ratio=alpha0_ratio, n_pw_iter=10, verbose=True
)
beta = directives.BetaEstimate_ByEig(beta0_ratio=1e-5, n_pw_iter=10)
betaIt = directives.PGI_BetaAlphaSchedule(
verbose=True,
coolingFactor=2.0,
progress=0.2,
)
targets = directives.MultiTargetMisfits(verbose=True)
petrodir = directives.PGI_UpdateParameters(update_gmm=False)
# Setup Inversion
inv = inversion.BaseInversion(
invProb,
directiveList=[alphas, scales, beta, petrodir, targets, betaIt, scaling_schedule],
)
mcluster_map = inv.run(minit)
# Inversion with no nonlinear mapping
reg_simple_no_map = regularization.PGI(
mesh=mesh,
gmmref=clfnomapping,
gmm=clfnomapping,
approx_gradient=True,
wiresmap=wires,
non_linear_relationships=False,
weights_list=[wr1, wr2],
)
opt = optimization.ProjectedGNCG(
maxIter=50,
tolX=1e-6,
cg_maxiter=100,
cg_rtol=1e-3,
lower=-10,
upper=10,
)
invProb = inverse_problem.BaseInvProblem(dmis, reg_simple_no_map, opt)
# directives
scales = directives.ScalingMultipleDataMisfits_ByEig(
chi0_ratio=np.r_[1.0, 1.0], verbose=True, n_pw_iter=10
)
scaling_schedule = directives.JointScalingSchedule(verbose=True)
alpha0_ratio = np.r_[100.0 * np.ones(2), 1, 1]
alphas = directives.AlphasSmoothEstimate_ByEig(
alpha0_ratio=alpha0_ratio, n_pw_iter=10, verbose=True
)
beta = directives.BetaEstimate_ByEig(beta0_ratio=1e-5, n_pw_iter=10)
betaIt = directives.PGI_BetaAlphaSchedule(
verbose=True,
coolingFactor=2.0,
progress=0.2,
)
targets = directives.MultiTargetMisfits(
chiSmall=1.0, TriggerSmall=True, TriggerTheta=False, verbose=True
)
petrodir = directives.PGI_UpdateParameters(update_gmm=False)
# Setup Inversion
inv = inversion.BaseInversion(
invProb,
directiveList=[alphas, scales, beta, petrodir, targets, betaIt, scaling_schedule],
)
mcluster_no_map = inv.run(minit)
# WeightedLeastSquares Inversion
reg1 = regularization.WeightedLeastSquares(
mesh, alpha_s=1.0, alpha_x=1.0, mapping=wires.m1, weights={"cell_weights": wr1}
)
reg2 = regularization.WeightedLeastSquares(
mesh, alpha_s=1.0, alpha_x=1.0, mapping=wires.m2, weights={"cell_weights": wr2}
)
reg = reg1 + reg2
opt = optimization.ProjectedGNCG(
maxIter=50,
tolX=1e-6,
cg_maxiter=100,
cg_rtol=1e-3,
lower=-10,
upper=10,
)
invProb = inverse_problem.BaseInvProblem(dmis, reg, opt)
# directives
alpha0_ratio = np.r_[1, 1, 1, 1]
alphas = directives.AlphasSmoothEstimate_ByEig(
alpha0_ratio=alpha0_ratio, n_pw_iter=10, verbose=True
)
scales = directives.ScalingMultipleDataMisfits_ByEig(
chi0_ratio=np.r_[1.0, 1.0], verbose=True, n_pw_iter=10
)
scaling_schedule = directives.JointScalingSchedule(verbose=True)
beta = directives.BetaEstimate_ByEig(beta0_ratio=1e-5, n_pw_iter=10)
beta_schedule = directives.BetaSchedule(coolingFactor=5.0, coolingRate=1)
targets = directives.MultiTargetMisfits(
TriggerSmall=False,
verbose=True,
)
# Setup Inversion
inv = inversion.BaseInversion(
invProb,
directiveList=[alphas, scales, beta, targets, beta_schedule, scaling_schedule],
)
mtik = inv.run(minit)
# Final Plot
fig, axes = plt.subplots(3, 4, figsize=(25, 15))
axes = axes.reshape(12)
left, width = 0.25, 0.5
bottom, height = 0.25, 0.5
right = left + width
top = bottom + height
axes[0].set_axis_off()
axes[0].text(
0.5 * (left + right),
0.5 * (bottom + top),
("Using true nonlinear\npetrophysical relationships"),
horizontalalignment="center",
verticalalignment="center",
fontsize=20,
color="black",
transform=axes[0].transAxes,
)
axes[1].plot(mesh.cell_centers_x, wires.m1 * mcluster_map, "b.-", ms=5, marker="v")
axes[1].plot(mesh.cell_centers_x, wires.m1 * m, "k--")
axes[1].set_title("Problem 1")
axes[1].legend(["Recovered Model", "True Model"], loc=1)
axes[1].set_xlabel("X")
axes[1].set_ylabel("Property 1")
axes[2].plot(mesh.cell_centers_x, wires.m2 * mcluster_map, "r.-", ms=5, marker="v")
axes[2].plot(mesh.cell_centers_x, wires.m2 * m, "k--")
axes[2].set_title("Problem 2")
axes[2].legend(["Recovered Model", "True Model"], loc=1)
axes[2].set_xlabel("X")
axes[2].set_ylabel("Property 2")
x, y = np.mgrid[-1:1:0.01, -4:2:0.01]
pos = np.empty(x.shape + (2,))
pos[:, :, 0] = x
pos[:, :, 1] = y
CS = axes[3].contour(
x,
y,
np.exp(clfmapping.score_samples(pos.reshape(-1, 2)).reshape(x.shape)),
100,
alpha=0.25,
cmap="viridis",
)
cs_proxy = mlines.Line2D([], [], label="True Petrophysical Distribution")
ps = axes[3].scatter(
wires.m1 * mcluster_map,
wires.m2 * mcluster_map,
marker="v",
label="Recovered model crossplot",
)
axes[3].set_title("Petrophysical Distribution")
axes[3].legend(handles=[cs_proxy, ps])
axes[3].set_xlabel("Property 1")
axes[3].set_ylabel("Property 2")
axes[4].set_axis_off()
axes[4].text(
0.5 * (left + right),
0.5 * (bottom + top),
("Using a pure\nGaussian distribution"),
horizontalalignment="center",
verticalalignment="center",
fontsize=20,
color="black",
transform=axes[4].transAxes,
)
axes[5].plot(mesh.cell_centers_x, wires.m1 * mcluster_no_map, "b.-", ms=5, marker="v")
axes[5].plot(mesh.cell_centers_x, wires.m1 * m, "k--")
axes[5].set_title("Problem 1")
axes[5].legend(["Recovered Model", "True Model"], loc=1)
axes[5].set_xlabel("X")
axes[5].set_ylabel("Property 1")
axes[6].plot(mesh.cell_centers_x, wires.m2 * mcluster_no_map, "r.-", ms=5, marker="v")
axes[6].plot(mesh.cell_centers_x, wires.m2 * m, "k--")
axes[6].set_title("Problem 2")
axes[6].legend(["Recovered Model", "True Model"], loc=1)
axes[6].set_xlabel("X")
axes[6].set_ylabel("Property 2")
CSF = axes[7].contour(
x,
y,
np.exp(clfmapping.score_samples(pos.reshape(-1, 2)).reshape(x.shape)),
100,
alpha=0.5,
label="True Petro. Distribution",
)
CS = axes[7].contour(
x,
y,
np.exp(clfnomapping.score_samples(pos.reshape(-1, 2)).reshape(x.shape)),
500,
cmap="viridis",
linestyles="--",
)
axes[7].scatter(
wires.m1 * mcluster_no_map,
wires.m2 * mcluster_no_map,
marker="v",
label="Recovered model crossplot",
)
cs_modeled_proxy = mlines.Line2D(
[], [], linestyle="--", label="Modeled Petro. Distribution"
)
axes[7].set_title("Petrophysical Distribution")
axes[7].legend(handles=[cs_proxy, cs_modeled_proxy, ps])
axes[7].set_xlabel("Property 1")
axes[7].set_ylabel("Property 2")
# Tikonov
axes[8].set_axis_off()
axes[8].text(
0.5 * (left + right),
0.5 * (bottom + top),
("Least-Squares\n~Using a single cluster"),
horizontalalignment="center",
verticalalignment="center",
fontsize=20,
color="black",
transform=axes[8].transAxes,
)
axes[9].plot(mesh.cell_centers_x, wires.m1 * mtik, "b.-", ms=5, marker="v")
axes[9].plot(mesh.cell_centers_x, wires.m1 * m, "k--")
axes[9].set_title("Problem 1")
axes[9].legend(["Recovered Model", "True Model"], loc=1)
axes[9].set_xlabel("X")
axes[9].set_ylabel("Property 1")
axes[10].plot(mesh.cell_centers_x, wires.m2 * mtik, "r.-", ms=5, marker="v")
axes[10].plot(mesh.cell_centers_x, wires.m2 * m, "k--")
axes[10].set_title("Problem 2")
axes[10].legend(["Recovered Model", "True Model"], loc=1)
axes[10].set_xlabel("X")
axes[10].set_ylabel("Property 2")
CS = axes[11].contour(
x,
y,
np.exp(clfmapping.score_samples(pos.reshape(-1, 2)).reshape(x.shape)),
100,
alpha=0.25,
cmap="viridis",
)
axes[11].scatter(wires.m1 * mtik, wires.m2 * mtik, marker="v")
axes[11].set_title("Petro Distribution")
axes[11].legend(handles=[cs_proxy, ps])
axes[11].set_xlabel("Property 1")
axes[11].set_ylabel("Property 2")
plt.subplots_adjust(wspace=0.3, hspace=0.3, top=0.85)
plt.show()
Total running time of the script: (0 minutes 31.954 seconds)
Estimated memory usage: 325 MB
